Circos¶
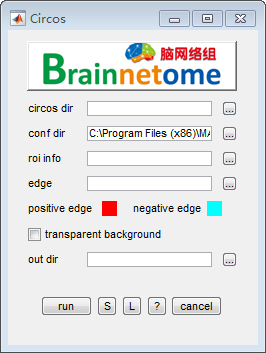
- circos dir: Path of unzipped circos folder.
/circos_path/bin - conf dir: Path of circos configure file.
- roi info: CSV table which defines sub-areas and lobes, there is an example in
brant-master/circos/brant_circos_3mm_273.csv - There should be at least four columns in the table:
- label: label of each sub-areas.
- module: to which lobe/module does the sub-area belong.
- index_module: order of the arranged module.
- index_node: order of sub-areas within one module.
Before using the function, please download and install circos 0.69 or higher.
- Buttons:
- S: Save parameters of the current panel to a
*.matfile. The*.matcan be further loaded for the panel or be used in a script processing. - L: Load parameters from
*.matfor the current panel. - ?: Help information.
- S: Save parameters of the current panel to a
- References:
- Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, et al. Circos: an information aesthetic for comparative genomics. Genome Res 2009; 19(9): 1639-45.
- Irimia A, Chambers MC, Torgerson CM, Van Horn JD. Circular representation of human cortical networks for subject and population-level connectomic visualization. Neuroimage 2012; 60(2): 1340-51.
- Fan L, Li H, Zhuo J, Zhang Y, Wang J, Chen L, et al. The Human Brainnetome Atlas: A New Brain Atlas Based on Connectional Architecture. Cereb Cortex 2016; 26(8): 3508-26.